Research in the lab can be split roughly into a number of connected areas
This page will be undergoing a major rewrite in Spring 2023 to reflect the evolution our research interests and what we have done over the years. Look for revamped pages comings soon!
Jan 2023 - First new 'Research Stories' page completed - 20 years of TRAP transporter :-)
If you are interested in working in the Thomas group then please get in touch through the Contact Us page. We are always looking for bright students who wish to study for a PhD or visiting students looking to spend some time in the UK working in the area of molecular microbiology.
Legacy material being updated and replaced in 2022 |
|
Structure and function of bacterial transporters
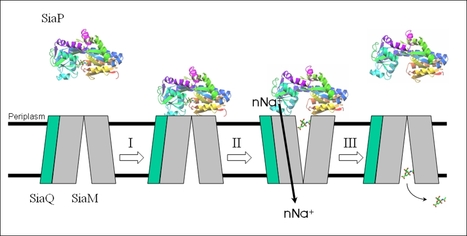
Model of TRAP transporter function. The SiaP structure was solved in York
The primary overarching interest of the lab is the discovery and characterisation of transporters in bacteria. For over a decade our main experimental system has been a family of transporters called the TRAP transporters, which stand for tripartite ATP-independent periplasmic transporters. These transporters are found all over the bacteria world and have important functions in many environments, primarily as organic acid transporters of high affinity.
Our main experimental systems are TRAP transporters that are specific for a sugar acid called sialic acid, which is important in the biology of the two bacteria from which we study it, namely Haemophilus influenzae and Vibrio cholerae. We mainly use biochemical and biophysical methods to study the function of the transporter combined with genetic and physiological experiments to understand their function in vivo. We were the first the solve the structure of a component of a TRAP transporter, the soluble substrate binding protein (SBP) component and have been able to fully reconstitute the transporter into proteoliposome for functional characterisation. Our main researchs aims at the moment are to understand how the SBP interacts with the membrane components of the transporter and how the system can couple the sodium-motive force to uptake of sialic acid. We are also trying to develop analogues of sialic acid which will inhibit the function of the transporter in collaboration with Dr. Paul Clark in the Department of Chemistry. Our work on the sialic acid TRAP transporter from V. cholerae was recently published in the Journal of Biological Chemistry and was picked up by the University and the BBSRC, our primary funders. We have recently started collaborating with Dr. Gregor Hageleuken at the University of Bonn, Germany, using PELDOR methods to investigate conformational dynamics of TRAP transporters.
As well as these TRAP transporters, we use similar methods to characterise transporters from many other families when relevant to our work, including related transporters for sialic acid and other substrates such as cell wall peptides. We have also recently had BBSRC funding to study a bacterial transporter involved in uptake of malodour precursors in Staphylococcal species in the human underarm.
Our main experimental systems are TRAP transporters that are specific for a sugar acid called sialic acid, which is important in the biology of the two bacteria from which we study it, namely Haemophilus influenzae and Vibrio cholerae. We mainly use biochemical and biophysical methods to study the function of the transporter combined with genetic and physiological experiments to understand their function in vivo. We were the first the solve the structure of a component of a TRAP transporter, the soluble substrate binding protein (SBP) component and have been able to fully reconstitute the transporter into proteoliposome for functional characterisation. Our main researchs aims at the moment are to understand how the SBP interacts with the membrane components of the transporter and how the system can couple the sodium-motive force to uptake of sialic acid. We are also trying to develop analogues of sialic acid which will inhibit the function of the transporter in collaboration with Dr. Paul Clark in the Department of Chemistry. Our work on the sialic acid TRAP transporter from V. cholerae was recently published in the Journal of Biological Chemistry and was picked up by the University and the BBSRC, our primary funders. We have recently started collaborating with Dr. Gregor Hageleuken at the University of Bonn, Germany, using PELDOR methods to investigate conformational dynamics of TRAP transporters.
As well as these TRAP transporters, we use similar methods to characterise transporters from many other families when relevant to our work, including related transporters for sialic acid and other substrates such as cell wall peptides. We have also recently had BBSRC funding to study a bacterial transporter involved in uptake of malodour precursors in Staphylococcal species in the human underarm.
Project DETOX: discovering how cells response to chemical toxicity and membrane damage (BBSRC IB Catalyse project)
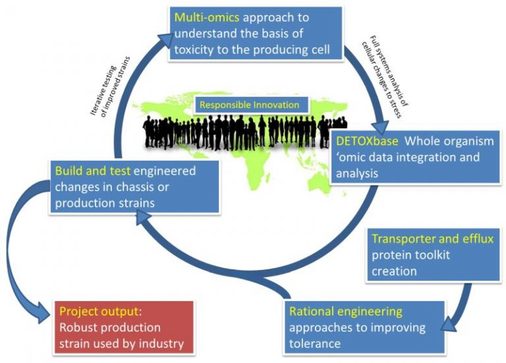
Product toxicity is one of the most serious barriers to commercialization of Industrial Biotechnology and Bio-Energy (IBBE). The DETOX project aims to provide the first ever systematic analysis of how chemicals poison bacterial cells. Our solutions will be used directly by UK IBBE business, growing the bio-manufacturing economy, reducing dependency on petrochemicals and creating a greener, more sustainable future (see Project WWW site for more details).
The project is a collaboration between academics at York, Nottingham, Sheffield, Cambridge and Exeter, and UK-based biotechnology companies Lucite, Green Biologics, Ingenza and the CPI, and is funded by the BBSRC. As the project develops we intend to extend our collaborations so please Contact Us to discuss the opportunities.
The project is a collaboration between academics at York, Nottingham, Sheffield, Cambridge and Exeter, and UK-based biotechnology companies Lucite, Green Biologics, Ingenza and the CPI, and is funded by the BBSRC. As the project develops we intend to extend our collaborations so please Contact Us to discuss the opportunities.
Systems biology of bacterial symbionts of insects
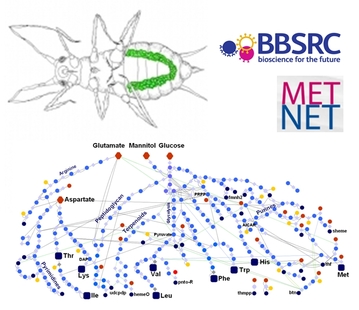
The aphid has an organ which contains Buchnera.
Many insects that live on restricted diets have an obligate need for a bacterial symbiont to provide scarce metabolites required for growth. We have been working with Prof. Angela Douglas at Cornell University on the symbiosis between the pea aphid and a bacterim Buchnera aphidicola, primarily as the bacterial genome is a subset of the model organism Escherichia coli K-12 which allows us to construct and analysis whole metabolic models of the symbiont. With BBSRC funding we have recently been able to understand much more about the symbiosis by studying genes and proteins enriched in the specialised cells that house the bacteria, called bacteriocytes. We have found a number of important extra roles for the symbiosis inlcuding recycling of purines from the aphid and other adaptations that feed the bacterium with its required metabolic precursors.
Sialic acid utilisation by bacteria
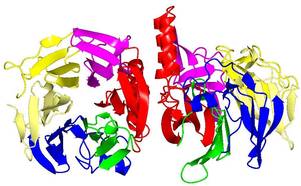
Structure of NanM solved in York
Form our work on the sialic acid TRAP transporter we have become more generally interested in how bacteria exploit sialic acid that is usually derived from their host. We have discovered an E. coli enzyme called NanM which is a sialic acid mutarotase. This enzyme allows bacteria to quickly convert the form of sialic acid that they release from host cell surfaces to the form that they transporter into their cells. We solved the structre of this protein in collaboration with Prof. Keith Wilson in the York Structural Biology Laboratory and also used NMR methods to show the mutarotase activity of the protein in collaboration with Prof. Jennifer Potts in the Department of Biology at York.
Escherichia coli bioinformatics
We have had a long-term interest in the model organism Escherichia coli K-12 and created a database, EchoBASE, that curates post-genomic information available about particular E. coli genes. This aims particularly at predicting function for uncharacterised E. coli genes. The most recent addition to this has been the EchoLocation tool, which integrates the outputs from many E. coli research tools to predict the subcellular location of E. coli gene products.
